Under the Hood #1: System Suitability Testing
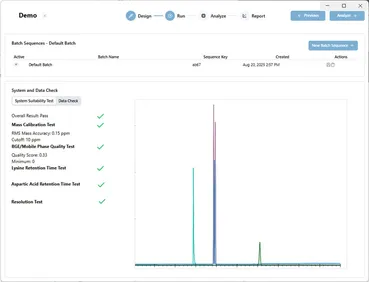
This post dives into how Move Analytical approaches one of the first steps in mass spectrometry metabolomics, system suitability testing. Specifically, we automate and simplify SST using software to move away from intuition and experience in evaluating the results of SSTs, in favor of simple tests with clear decision criteria.
Motivation
The desired outcome of a system suitability test (SST), at the risk of being obvious, is to determine if your “system” (analyzer) is “suitable” to run samples, typically on the day of the measurement. In the context of a mass spectrometry-based SST, the broad goal is to make sure you don’t waste time running samples only to find out the data is unsatisfactory because your MS was in poor operating condition. Running an SST mitigates the risk of MS contamination, improper calibration, and a host of potential separation failure modes.
For simple analytical tests, the SST can be very straightforward. Imagine a hypothetical SST for an analytical balance, which makes a single measurement (mass); such a test may consist of using a standard of known mass and verifying that the system’s measurement is correct. For mass spectrometry measurements, SST is much more complicated even for single analytes, and for ‘omics methods there are so many analytes and variables that performing and interpreting the SST can be intimidating if not downright paralyzing.
Establishing appropriate SSTs is required as part of the method validation process for regulated bioanalysis. Performing SST in metabolomics applications has also become commonplace (Mosley JD 2024). However, publications on SST in metabolomics and proteomics have mostly focused on the wide array of metrics that can be tracked (Rudnick PA, 2010), on tools for establishing and tracking results with reference-ranges using Westgard rules (Bereman MA, 2014), or on the overall utilization of SST in the community (Mosley JD et al. 2024). What they have mostly failed to do thus far is to guide practitioners to clear decision rules that can support direct “go/no go” decisions that do not rely on intuition and experience or difficult-to-establish reference ranges.
How Can It Be Done Differently?
We believe that proper SST in metabolomics can be made more accessible to new or inexperienced users, as well as more reproducibly implemented in experienced labs, by reducing the number of measurements and removing the need for fuzzy decision rules. A carefully designed test should be tailored to the specific task of metabolomics, easy to run, and easy to interpret and act on.
Getting the right answer is the most important job of an SST, so we have carefully chosen metrics that reveal the critical operating characteristics of the system. But we also believe that using the fewest metrics that give the right answer (i.e. ignoring things you could measure, on purpose) has multiple benefits: it makes for a simpler test that is easier to deploy, and it is less likely to give false-positive failures. Making the test simple means that users are more likely to do it, and decreasing false-positives means that users are less likely to ignore a failure when it happens. Both of these benefits should result in increased adoption and adherence to the SST results for both experienced and inexperienced users.
Additionally, we have chosen hard cutoffs instead of reference ranges wherever possible in designing SSTs. While reference ranges can be more sensitive, they are more difficult to establish (especially across different laboratories) and can require complex trend monitoring. Again, we believe simplification can play a significant role in improved results across the community.
Finally, all readouts need to be interpretable and actionable. As an example, a clinical LC-MS/MS test for thyroid hormones T3/rT3 uses a cutoff for chromatographic resolution in system suitability because these analytes cannot be differentiated by tandem MS, and thus chromatographic selectivity is critical for accurate quantification (private communication, Chris Shuford, Labcorp). In that case, insufficient resolution in the separation means inaccurate quantification. If that simple test fails, it implies a clear action: to troubleshoot the chromatography (usually column or mobile phase).
Practical Application
In our metabolomics kits and software, we have used deep domain knowledge and thousands of sample analyses to simplify the SST sample and test as much as possible. For MoveKit™ CE, the test mixture contains 17 amino acids, and all SST metrics are calculated from the interpretation of 5 analytes. If a test result falls outside the established thresholds for any of the SST test metrics, we guide the user to specific recommended corrective actions. We utilize the following tests:
-
Mass Accuracy. Compounds in the SST mix across the mass range are evaluated for their mass accuracy, which is important for compound ID and quantification. The corrective action is typically to calibrate the mass spectrometer.
-
BGE/Mobile Phase Quality. Analyte elution position relative to one another is critical to using time indexing for reliable compound quantification against a standards library. In MoveKit™ CE we calculate a simple quality metric using relative migration time of three compounds which are known to be highly sensitive and differentially affected by background electrolyte (BGE) pH. The corrective action is to check BGE composition and expiration, and reprime the fluidics.
-
Analyte Migration/Retention. We use the first and last eluting compound to ensure detection of metabolites across the expected elution window. This is primarily to check that the correct column/chip and SST sample is being used.
-
Separation Resolution. The resolution of a critical pair of isomeric compounds is used to ensure that they are sufficiently separated for accurate metabolite quantification of all analytes in the library, without unwanted interference. Corrective actions here are to reprime the system and if a repeat failure occurs check the age/expiration of the chip and BGE and replace as necessary.
What About All My Other Favorite Metrics?
Every experienced mass spectrometrist has their own favorite system suitability metrics; their own favorite “canary” metrics for their assay, if you will. We are not discounting this wealth of knowledge. We are only trying to simplify the measurements to make the decision more accessible and less subjective. If you’re one of these MS experts, you might have one question right away: we are only using 5 out of 17 molecules in the SST to calculate the pass/fail criteria, so why have extras? The full story is that the number of molecules in the mixture reflects our active research process, where we began development using an off-the-shelf commercial product and then carefully selected the minimal required set to make the correct GO/NO GO decisions with automation. We continued using the same qualified material as our SST mixture instead of making a new one.
Admittedly, there are lots of other items which others have highlighted for SST performance which we are ignoring. For instance, what about sensitivity? This is a great example of another concerted effort to use the fewest metrics that will give the right answer. In an effort to simplify wherever possible, we made the concentration of the SST sample low enough so that if it passes the other metrics, sensitivity will be good enough for the overall assay. By avoiding the need for an explicit sensitivity test, we avoid questions around intensity cutoffs and subjective judgement calls on degrees of sensitivity drift which are likely irrelevant for assay performance.
Of course, we are always looking for ways to improve our product so if we find that we currently exclude a measurement that is important to spot an SST failure in a specific edge or use-case, we will update our SST test and protocol. We remain dedicated to listening to, and working with, our customers to continually iterate our products and ensure the automated approaches don’t miss important SST failures.
Closing Thoughts
SST is just the start of our automation journey, as it should be the start of any quality analytical measurement. By using the fewest metrics for SST which give the correct answer, actionable cutoffs, and automated interpretation to a GO/NO GO decision, we seek to eliminate at least one pain/decision point in the analytical lab. We hope application of SST in MoveKit™ will help users to embrace our mission of Making Metabolomics Easy while maintaining the high-quality results we all expect, and encourage the metabolomics community to consider these concepts when they look to build their own SSTs.
References
Bereman MS, Johnson R, Bollinger J, et al. Implementation of statistical process control for proteomic experiments via LC MS/MS. J Am Soc Mass Spectrom. 2014 Apr;25(4):581-7.
Mosley JD, Schock TB, Beecher CW, et al. Establishing a framework for best practices for quality assurance and quality control in untargeted metabolomics. Metabolomics 2024 Feb 12;20(2):20.
Rudnick PA , Clauser KR, Kilpatrick LE et al. Performance metrics for liquid chromatography-tandem mass spectrometry systems in proteomics analyses. Mol Cell Proteomics. 2010 Feb;9(2):225-41.
About the Authors
J. Will Thompson is a Founding Partner and serves as Operations Lead at Move Analytical LLC. James Campbell is a Founding Partner and serves as Software and AI Lead at Move Analytical.
